
May miss filtering capacities (only minimum peak distance with the delta parameter). The algorithm was written by sixtenbe based on the previous work of endolith and Eli Billauer.Įasy to setup as it comes in a single source file, but the lookahead parameter make it hard to use on low-sampled signals or short samples. # with for each peak a tuple of (indexes, values). # peakdetect returns two lists, respectively positive and negative peaks, Peaks = peakdetect.peakdetect(np.array(vector), lookahead=2, delta=2) Print('Detect peaks with distance filters.') This algorithm can be used as an equivalent of the MatLab findpeaks and will give easily give consistent results if you only need minimal distance and height filtering. The function has an interface very similar and consistent results with the MatLab Signal Processing Toolbox findpeaks, yet with less complete filtering and tuning support. This algorithm comes from a notebook written by Marcos Duarte and is pretty trivial to use. Indexes = detect_tect_peaks(vector, mph=7, mpd=2) _peaks searches for peaks (local maxima) based on simple value comparison of neighbouring samples and returns those peaks whose properties match optionally specified conditions (minimum and / or maximum) for their height, prominence, width, threshold and distance to each other. This function was added to SciPy in version 1.1.0 and is comparable to findpeaks provided in Matlab's Signal Processing Toolbox. Indexes, _ = _peaks(vector, height=7, distance=2.1) Print('Detect peaks with minimum height and distance filters.') See also related functions argrelmin and argrelmax. The filtering behavior is customizable through the comparator parameter, whichĬan make it a good choice for building your own filtering algorithm over it. It includes an order parameter that can serve as a kind of minimum distance filter. Its usage is really trivial,īut it misses out of the box filtering capacities. New peak detection algorithm from Scipy since version 0.11.0. Print('Detect peaks with order (distance) filter.') 
However the wavelet approach can make it a good choice on noisy data. This is less trivial and direct than other algorithms. This function requires to understand wavelets to be properly used. Its name appears to make it an obvious choice (when you already work with Scipy), but it may actually not be, as it uses a wavelet convolution approach. The first historical peak detection algorithm from the Scipy signal processing package. Indexes = _peaks_cwt(vector, np.arange(1, 4), 
Print('Detect peaks without any filters.') Does it require extra dependency? Does is it easy to make it run on a fresh box? You may want the function to work natively with Numpy arrays or may search something similar to other platform algorithms, like the MatLab findpeaks. When you're selecting an algorithm, you might consider: time) at fixed x- and y-locations using the default behaviour of the FINDPEAKS function.Overview of the peaks dectection algorithms available in Python. The second cell searches values across the third dimension (i.e. I've noted the values pertinent to your needs. 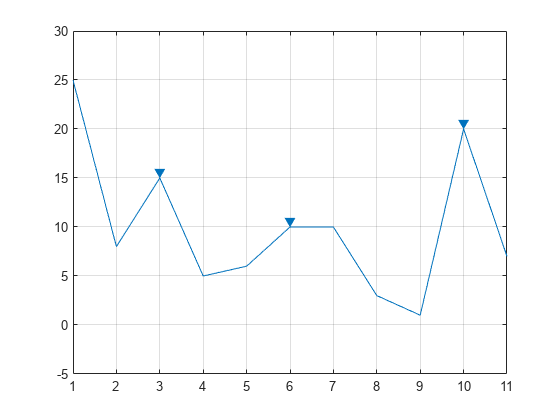
the %% header) creates the 3-dimensional matrix that has been suggested. WorkingSet = squeeze(a3DMatrix(xIndex, yIndex, :)) Using a 3-dimensional matrix also reduces the mental effort of jumping back-and-forth between cell-arrays and matrices.Ĭonsider, %% Concatenate Cell-arrays to Create 3D MatrixĪ3DMatrix = nan(3, 3, 3) % nan(101, 151, 476)Ī3DMatrix(:, :, cellArrayIndex) = surfaceArray įprintf(1, '\n(x, y) = (%d, %d)\n', xIndex, yIndex) the same (x, y) pair across the cells) and the FINDPEAKS function only works on a vector, it would be easier produce a 3-dimensional matrix. Since you want to find the peaks at each x- and y-value along the time axis (i.e.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
